Once more time, with feeling! Toward reproducible computational science
My scientific education was committed at the hands of physicists. And though I've moved on from academic physics, I've taken bits of it with me. In MIT's infamous Junior Lab, all students were assigned lab notebooks, which we used to document our progress reproducing significant experiments in modern physics. It was hard. I made a lot of mistakes. But my professors told me that instead of erasing the mistakes, I should strike them out with a single line, leaving them forever legible. Because mistakes are part of science (or any other human endeavor). So when mistakes happen, the important thing is to understand and document them. I still have my notebooks, and I can look back and see exactly what I did in those experiments, mistakes and all. And that's the point. You can go back and recreate exactly what I did, avoid the mistakes I caught, and identify any I might have missed. Ideally, science is supposed to be reproducible. In current practice though, most research is never replicated, and when it is, the results are very often not reproducible. I'm particularly concerned with reproducibility in the emerging field of computational social science, which relies so heavily on software. Because as everyone knows, software kind of sucks. So here are a few of the tricks I've been using as a researcher to try to make my work a little more reproducible.
Databases
When I'm doing anything complicated with large amounts of data, I often like to use a database. Databases are great at searching and sorting through large amounts of data quickly and making the best use of the available hardware, far better than anything I could write myself in most cases. They've also been thoroughly tested. It used to be that relational databases were the only option. Relational databases allow you to link different types of data using a query language (usually SQL) to create complicated queries. A query might translate to something like "show me every movie in which Kevin Bacon does not appear, but an actor who has appeared in another movie with Kevin Bacon does." A lot of the work is done by the database. What's more, most relational databases guarantee a set of properties called ACID. Generally speaking, ACID means that even if you're accessing a database from several threads or processes, it won't (for example) change the data halfway through a query.
In recent years, NoSQL databases (key-value stores, document stores, etc.) have become a popular alternative to relational databases. They're simple and fast, so it's easy to see why they're popular. But their simplicity means your code needs to do more of the work, and that means you have more to test, debug, and document. And the performance is usually achieved by dropping some of the ACID requirements, meaning that in some cases data might change in the middle of a calculation or just plain disappear. That's fine if you're writing a search engine for cat gifs, but not if you're trying to do verifiably correct and reproducible calculations. And the more data you're working with, the more likely one of these problems is to pop up. So when I use a database for scientific work, I currently prefer to stick with relational databases.
Unit tests
Some bugs are harder to find than others. If you have a syntax error, your compiler or interpreter will tell you right away. But if you have a logic error, like a plus where you meant to put a minus, your program will run fine, it'll just give you the wrong output. This is particularly dangerous in research, where by definition you don't know what the output should be. So how do you check for these kinds of errors? One option is to look at the output and see if it makes sense. But this approach opens the door to confirmation bias. You'll only catch the bugs that make the output look less like what you expect. Any bugs that make the output look more like what you expect to see will go unnoticed.
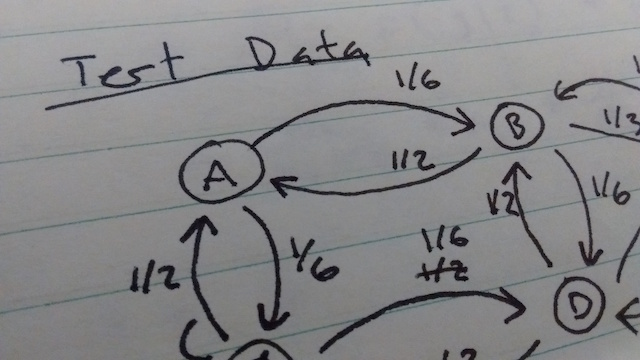
So what's a researcher to do? This is where unit tests come in. Unit tests are common in the world of software engineering, but they haven't caught on in scientific computing yet. The idea is to divide your code into small chunks, and test each part individually. The tests are programs themselves, so they're easy to re-run if you change your code. To do this in a research context, I like to compare the output of each chunk to a hand calculation. I start by creating a data set small enough to analyze by hand, but large enough to capture important features, and writing it down in my lab notebook. Then for each stage of my processing pipeline, I calculate what the input and output will be for that data set by hand and write that down in my lab notebook. Then I write unit tests to verify that my code gives the right output for the test data. It almost always doesn't the first time I run it, but more often than not it's a problem with my hand calculation. You can check out an example here. A nice side effect of doing unit tests is that it gives you confidence in your code, which means you can devote all of your attention to interpreting results, rather than second guessing your code.
Version control
Version control tools like git are becoming more common in scientific computing. On top of making it easy to roll back changes that break your code, they also make it possible to keep track exactly what the code looked like when an analysis was run. That makes it possible to check out an old version of code and re-run an analysis exactly. Now that's how you reproducible! One caveat here: in order to keep an accurate record of the code that was run, you have to make sure all changes have been committed and that the revision id is recorded somewhere.
Logging
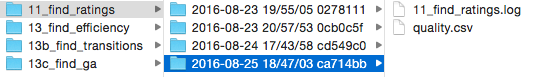
Finally, logging the process of analysis scripts makes it a lot easier to know exactly what your code did, especially years after the fact. In order to help match my results with the version of code that produced them, I wrote a small logging helper script that automatically opens a standard python log in a directory based on the experiment name, timestamp, and current git hash. It also makes it easy to create output files in that directory. Using a script like this makes it easy to know when a script was run, exactly what version of the code it used, and what happened during the script execution.
As for the specific logging messages, there are a few things I always try to include. First, I always have a message for when the script starts and completes successfully. I also wrap all of my scripts in a try/except block and log any unhandled exceptions. I also like to log progress, so that if the script crashes, I can know where to look for the error and where to restart it. Logging progress also makes it easier to estimate how long a script will take to finish.
Using all of these techniques has definitely made my code easier to follow and retrace, but there's still so much more that we can do to make research truly open and reproducible. Has anyone reading this tried anything similar? Or are there things I've left out? I'd love to hear what other people are up to along these lines.
